If you want to be a future molecular biologist or may be interested in tiny little particles inside your body that determines whether you will be the tallest/shortest person in your school or why your eye color is black not green, then surely I will recommend you to have a look in this following article.
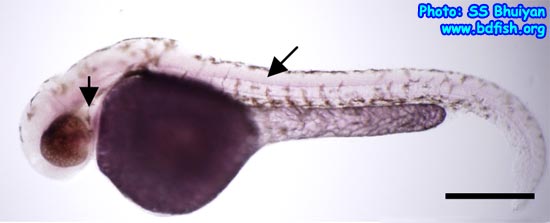
The word ‘in situ’ came from the original Latin phrase which means in place or position. In situ hybridization (ISH) is a technique which is used by molecular biologists to detect the position or localizing of a gene in cell, tissue or in the chromosome. This can be easily detected by staining (colorization) with gene specific antibody (Fig. 1 a-b).

The technique for gene detection was time consuming as well as costly for the last few decades and thus restricted scientists to study deeply about the expressional and functional study of gene. If a new gene is discovered, scientists try to discover first where that gene is working (expressional study) and what is the probable function (functional study) of that particular gene. To find the answers, ISH was first described independently by the scientists Gall and Pardue (1969), Buongiorno-Nardelli and Amaldi (1969), and John et al. (1969). Since then many variations of ISH methods have been developed.
During the last decade, ISH has undergone many refinements and has become an important tool in basic scientific research of molecular biology. Between January 1994 and July 1995, more than 4,000 articles had been published using this technique. These rapid advances have made ISH as one of the most important tools of molecular biology study.
The technique for in situ hybridization is quite simple and easy though time consuming. For in situ hybridization, first the newly discovered gene’s Deoxy Ribo-nucleic Acids (DNAs) sequence is collected from the database such as popular gene bank databases (eg. http://uswest.ensembl.org, or The DDBJ/EMBL/GeneBank database etc).
Suppose the newly discovered gene is found in human and we want to test whether the same gene is functioning in fish or not, we collect the double stranded DNA sequence of that target gene from the gene database and we make short (about 20 base nucleic acid containing) single stranded DNA sequence which is called primer.
We can design primers from many available primes designing tool that can be found in the internet for free (eg.Oligocalculator: http://www.pitt.edu/~rsup/OligoCalc.html).
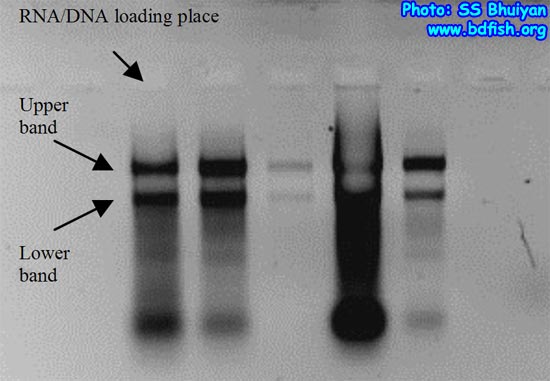
From the target fish we extract total Ribo-Nucleic Acids (RNA) which is single stranded following any reliable manuals as there are many different ways for RNA extraction from fish (eg. http://www.ambion.com/techlib/basics/rnaisol/index.html) and we can check the success of RNA extraction by a process called electrophoresis, a process explained at later phase (Fig. 2).
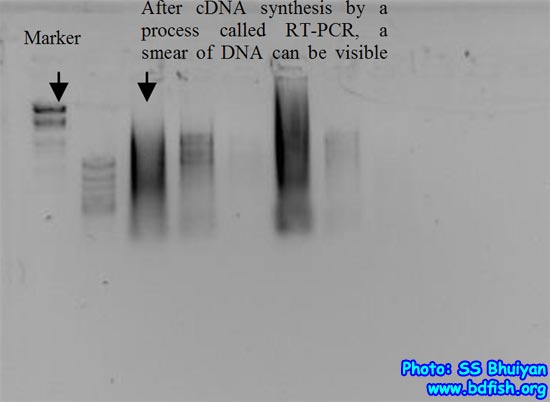
After that, we use a technique called Reverse Transcriptase Polymerase Chain Reaction (RT-PCR) which can generates from a single stranded RNA to double stranded DNA, called complementary DNA strands (cDNAs), usually appear a smear if we check the result of cDNA making success by gel electrophoresis (fig. 3).
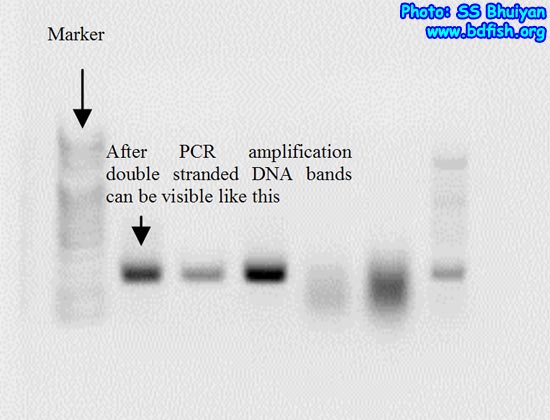
Now we come back to the primers that was designed previously and make copy specific (which is called amplification) of the cDNAs to double stranded DNA (Fig. 4) using the previously designed primer. This process is widely known as Polymerase Chain Reaction or popularly known as PCR. After PCR is carried out we can check the success of the result by a process called electrophoresis in a machine called electrophoresis apparatus (Fig. 5-6).


Continued…
Basic term and apparatus used in this article:
- DNA and RNA:
Both DNA and RNA are macromolecules identified by Watson and Crick in the late 1950s’ of which DNA is double helical (double stranded) and RNA contain a single helix (Single stranded).
- Gene:
Each cell in the human body contains about 25,000 to 35,000 genes, which carry information that go toward determining traits. Traits are characteristics you inherit from your parents; Genes are found in all animals and plants.
- cDNA:
Complementary DNA (cDNA) is a doublestranded DNA version of an mRNA molecule.
- Gel Electrophoresis:
Electrophoresis means migration of suspended particles such as protein macromolecules, under the influence of an electric field.
- Electrophoresis apparatus:
It is a buffer-filled box used for the purpose of electrophoresis. In this machine, a gel loaded with DNA/RNA is placed and electrical field (positive and negatively charged end) is applied via the power supply. As DNA is negatively charged so it migrates to the positive end.
- Marker:
A marker is a solution of DNA molecules with different lengths used in gel electrophoresis. It is applied to a gel as a reference to estimate the size of unknown DNA molecules.
References:
- Buongioino N, M., and F. Amaldi. 1970. Autoradiography detection of molecular hybrids between rRNA and DNA present in tissue sections. Nature (Lond.).225:946.
- Gall, J.G., and L. Pardue. 1969. Formation and detection of RNA-DNA hybrid molecules in cytological preparations. Prec.Natl.Acad.Scz.USA.63:378.
- John, H A., L. Birmstiel, and K .W. Jones.1969. RNA-DNA hybrids at the cytological level. Nature (Lond.).223:582.
Visited 1,164 times, 1 visits today | Have any fisheries relevant question?
